Heinrich Mallison’s photo-rific dinosaurpalaeo blog has the first of what might, if the Gods of the Freezers remain kind, be a series of posts on our dissections of some of the verrrrrry same giraffe limbs featured earlier on this blog. Have a brush with greatness- see the giraffe legs in deconstruction! For free! What more fun could you possibly have (legally)?
Posts Tagged ‘giraffe’
Head over to dinosaurpalaeo for Mammal Monday 17 and a Giraffe Dissection!
Posted in Frozen Mammals, tagged anatomy, buddies, dissection, giraffe on April 2, 2012| 4 Comments »
Awesome giraffe dissection photo!
Posted in Frozen Mammals, tagged anatomy, dissection, giraffe on March 6, 2012| 4 Comments »
William Pérez from the Veterinary Anatomy Facebook page sent me a link to this stunning image of a giraffe hindlimb dissection– wowza!
Scanner’s eye view of giraffe leg
Posted in Frozen Mammals, tagged CT, giraffe on March 6, 2012| 1 Comment »

This is why we get scan artifacts from giant specimens. It fits, but only just. The x-ray beams are getting scattered from being too close to the x-ray detectors (ring around the specimen), creating noise in the images. The red lines on the specimen are from a laser, used to align it properly within the gantry of the scanner. The femoral head (hip joint) is visible as the pale white thingy, down on the bottom right.
3D rendering of CT data for giraffe hindlimb is easy!
Posted in Frozen Mammals, tagged CT, giraffe, modelling, reconstruction on March 6, 2012| 2 Comments »
We use Mimics software, which is pricey but sooooo easy to do stuff like this. Here I’ve just had a very quick pass at reconstructing the distal hindlimb (limb except the thigh, which had bad scan artifacts). With more effort, I could remove the pesky artifacts around the knee and ankle (2 uppermost joints), although some of those are unavoidable because the leg was too damn big for the CT scanner gantry (70cm diameter; leg was around 60cm across at largest).
So how long did this process take? The hardest part was moving the leg around and positioning it on the CT table. Then the scan (608 cross-sectional slices) took about 15 minutes to do, then 15 more minutes to transfer over to my PC. Loading the DICOM slices of data and making a movie (previous post) took 5 minutes, and then making this 3D reconstruction movie took just another 5 minutes, although I waited a few days because I was busy.
So, when operating at peak efficiency, we can obtain decent 3D models from frozen specimens in less than an hour. This is but one example of how modern technology, especially X-ray computed tomography and computer hardware/graphics software, have massively transformed any research that deals with anatomy. When I was doing my PhD back in the 90’s, this would have been a much more time-intensive procedure (probably weeks of work, and difficulties getting CT access); in the 80’s it would basically have been impossible.
In the future, now, we’ll be using these models (once cleaned up a bit) with data from dissection to model how the limbs work in a real giraffe. More about that later. The Giraffe-A-Thon is over for now. I hope you enjoyed it! More of the same to come on this blog!
Giraffe right hindlimb: what a CT scan shows you
Posted in Frozen Mammals, tagged CT, giraffe on March 5, 2012| 1 Comment »
Giraffe limb from previous post, now shown via movie of DICOM (CT image data) files. Axial slices every 2.5mm, from toes to knee/stifle. Darker areas are lower density (black is air); white is very dense– bone, artefact, metal, Santorum, etc.
Right hindlimb of giraffe ready for CT scan!
Posted in Frozen Mammals, tagged CT, giraffe on March 3, 2012| 8 Comments »
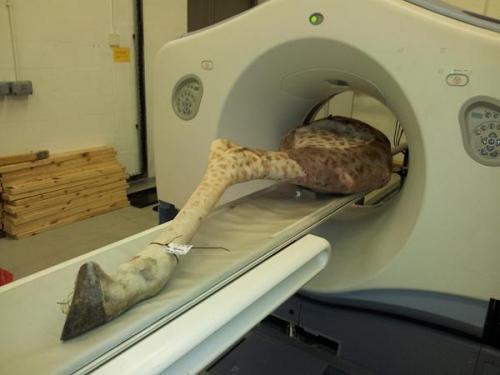
I’m preparing to do anatomically-realistic computer modelling of giraffe locomotor mechanics with some colleagues. To do that, we of course need the 3D anatomy of bones, muscles and tendons, for which CT can be pretty useful. Here, we put our first frozen leg through the motions. It was a 3 person job to lift the sucker, but the CT bed managed to move it through the scanner with minimal hiccups. Inside the ring that the upper end of the leg is lined up with are 8 x-ray detectors, so 8 CT slices can be imaged at once, speeding the procedure.
This specimen died in a UK zoo recently, apparently from trauma (falling?), which we’re trying to help them figure out in the course of our scans and future dissections. We often provide a pretty detailed postmortem service in return for being given cadavers, since we are a vet school with a lot of expertise in pathology and anatomy. Also, we have been describing the kinds of pathologies we observe along the way, because terribly little is known about some diseases/injuries in non-domestic animals, so there is plenty we can contribute to the scientific literature as a result. We’re also interested in documenting how pathologies in wild vs. captive animals differ (if at all).

